Locus Search¶
A Locus search enables the user to upload any sequence and get loci identified and alleles assigned for any scheme in the database. Thus, is can allow the identification of alleles for 7 gene MLST from a fasta file of allele sequences. In addition you could upload the sequence of an operon, phage or plasmid you are interested in and have that tagged with wgMLST loci. Furthermore loci in other strains in Enterobase can than be compared to the sequence that you uploaded.
Creating A Locus Search¶
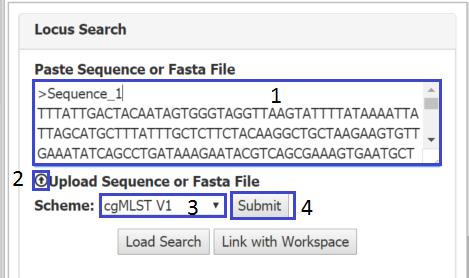
The Locus Search page can be reached from left hand menu under Tasks > Locus Search. In the top left hand panel textbox (1) paste a sequence of fasta file (the fasta file can contain multiple sequences). Or a sequence/fasta file can be uploaded using the button below the text box(2) . Make sure you select the scheme you require (3) and press submit. A dialog will appear allowing you to set a name and location for your locus search. If the search is taking a while, you can leave the page, return later to load the saved search (5).
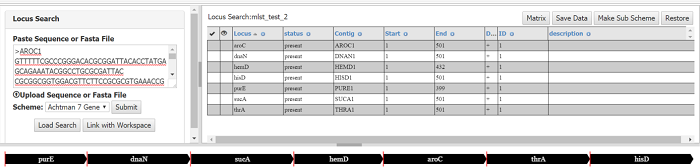
When a locus search is complete (or a previous search loaded) , a table will appear in the top right panel showing all the loci found in the input sequences and their position. Allele IDs are also given, however if a locus is discovered which contains a new allele it will not be assigned a new ID, since the validity and quality of the uploaded sequence cannot be ascertained. A Graphic representation is also given. Any contigs are just stitched together (in the order of the sequence) and boundaries between contigs are shown as red lines.
Link With Workspace¶
This facility allows you to examine the loci present on the uploaded sequence for a given group of strains
Clicking the ‘Link With Workspace’ button (5) will display a dialog showing all the workspaces/trees that you have access to ([Workspaces/trees](workspaces) can be created from the main search page of the database of interest). Select a tree/workspace that you want to link to the locus search and press load. In the dialog, clicking on a workspace will reveal any trees that are are part of the workspace. Trees or workspaces can be chosen, but if a tree is chosen, the genomes will be ordered by their position in the tree. At the moment only SNP Trees can be viewed- but it hoped that MS Trees will be displayed in the near future. When linking a workspace for the first time, it may take a few seconds to calculate, but results are cached , so subsequent analysis should be quicker.
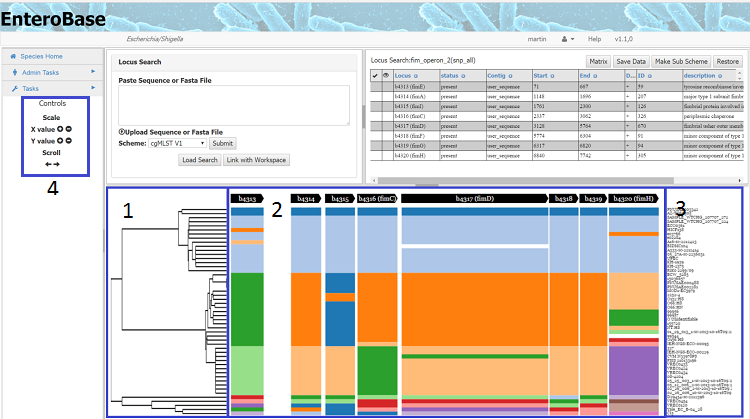
Once the workspace has been linked, a graph will appear below the loci, the left hand section of which (1) shows the tree, but will be missing if a workspace is chosen. The right hand section shows the strain names(3) and the middle section the map showing the presence/absence of loci in each strain, loci are coloured by allele number. . Mouse over a loci to find the name of the locus, the allele number and the strain name. N.B. the position of loci on the map does not necessarily reflect their position in that specific genome (to obtain this information see below).
The Controls for the graph are in the left hand menu panel (4). The x and y axis can be increased/decreased independently, but by using the mouse wheel allows zooming in and out is also possible. This panel also allows left and right scrolling, but scrolling in all directions can be achieved by dragging on the graph itself.
Obtaining More Information on Loci¶
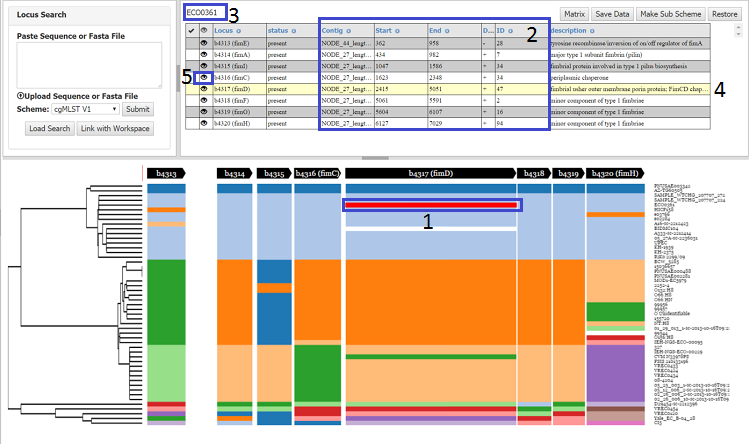
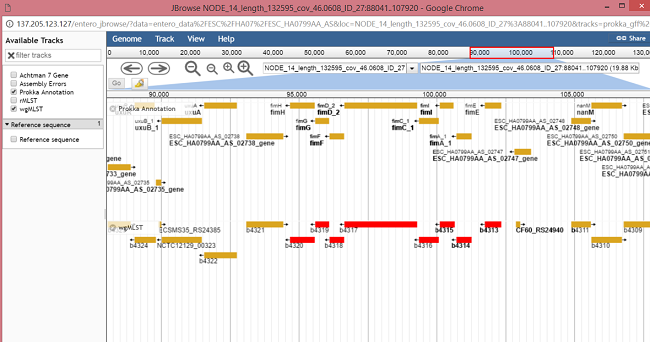
Clicking on a locus will highlight all loci in that particular block (1) and strain specific information (genomic location ,allele ID etc.) for all loci in the block will be shown in the table (4), with the name of strain being shown above the table (3). The actual locus clicked will be highlighted in the table (2). Clicking on the eye icon (5) for a locus will open up a genome viewer (JBrowse), which shows the position of that locus in the genome.
All loci in the selected block will be red in the wgMLST track. The other track shown by default displays all the genes in the prokka annotation, which may contain genes that are not in the wgMLST scheme and will contain more information on each gene (locus). Depending on the type of genome, other tracks (selected from the left hand panel) will be available e.g Assembly Errors, GenBank Annotation, other schemes etc. If the genome has not been viewed before it will have to be formatted which will take a few seconds, so please be patient.
Exporting Data¶
The ‘Matrix’ button above the table will download a matrix containing allele IDs for loci (columns) in all the strains (rows). Absent loci will have an allele ID of 0. ‘Save Data’ will download all the data present in the table
Creating A Sub-scheme¶
A sub-scheme ([Custom View](User-Defined Content)) Can be created by checking the loci that you want in the main table and then pressing the ‘Make Sub Scheme’ button. A dialog will then appear which allows you to specify the name and location of the sub scheme (custom view), which can be viewed in the main search page.