GrapeTree Reference Manual¶
This page explains each of the features in GrapeTree.
Interaction With Enterobase¶
- Load Selected: Any strains selected in the tree will be loaded into an Enterobase workspace, opening a new browser window if necessary.
- Highlight Checked: Any strains that are selected (checked) in Enterobase will be highlighted (large yellow halo) in the GrapeTree. However, it is often simpler to select them in the GrapeTree metadata table (right click, Show metadata table), which synchronises imediately with the tree.
- Import Fields: Shows a dialog box which allows the selection of experimental fields and custom fields (columns) to be imported into the tree. The last field that is imported is immediately used for colour-coding of the nodes and for the figure legend.
- Save:Saves a local copy of the tree layout and any metadata, which will be restored when this GrapeTree is opened again from EnteoBase. Saving changed metadata in the metadata table does not update metadata in Enterobase. However, saving changed data in user-defined custom fields will updated those fields in Enterobase.
- Update: Will update the any metadata or user-defined fields within the GrapeTree metadata tree with the current version of the metadata and user-defined fields in Enterobase.
- Info: Shows information about the tree such as the parameters used for construction, number of strains, last modified etc.
Input to Standalone GrapeTree¶
To get started, Drag and drop one of the following types of files into the browser window or click Load Files
Trees or Profile Data
- Phylogenetic trees: GrapeTree accepts trees in either Nexus or Newick (nwk) format.
- Allelic Profiles: These are tab-delimited text files with columns of alleles and rows of strains. See Tutorial 1B: Basic Usage of GrapeTree (Stand-alone). A header line is required, and column in which headers start with a ‘#’ are ignored. Importing allelic profiles does not work with the demo GrapeTree hosted on GitHub. The profile data need to be any alphanumeric text. Missing data can be indicated by - or 0.
- GrapeTree Format(.json): Previous saved sessions in GrapeTree can be restored by loading a saved JSON version, which will contain the previous tree and metadata.
Metadata
- Metadata files The first column (column header ID) must contain a unique ID which is also present in the Newick/Nexus tree file.
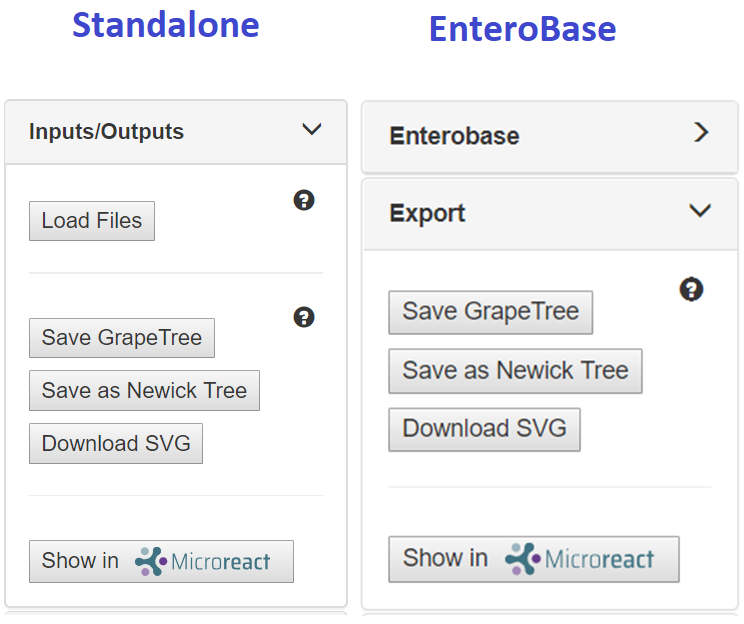
Outputs¶
- Save GrapeTree: Save a local copy of the current session in GrapeTree (JSON) format. This includes the current display form of the tree data plus any metadata.
- Save as Newick Tree: Save the tree as a Newick (nwk) file, which contains tree topology, branch lengths and tip names. Compatible with most tree visualisation tools.
- Download SVG: Save the tree as a Scalar Vector Graphic (SVG) file, a vector image that can be loaded into image publishing software such as Inkscape or Abobe llustrator.
- Transmit phylogeny and metadata to MicroReact
MLST trees and SNP projects can be transferred to MicroReact via the GrapeTree interface. Firstly, adjust the tree to contain only the desired branches by selecting them (shift Click) and right-clicking on the screen (Show selected subtrees). Ensure that all metadata that might be useful within MicroReact are included in the GrapeTree (EnteroBaseImport Fields or upload a modified metadata table into standalone GrapoeTree). Choose the node color-coding in GrapeTree (right click on Figure Legend. Choose from the DropDown List titled Colour by). Send the data to MicroReact using ExportShow in MicroReact in the left pane.
Details on transferring data to MicroReact¶
MicroReact locate genomes on a map using GPS coordinates. EnteroBase will transfer GPS co-ordinates if they are included in the metadata. Standalone GrapeTree will recognise columns with headers labelled Latitude and Longitude and which contain decimal data as GPS co-ordinated. Where geographical locations are available instead of GPS co-ordinates, GrapeTree transmits the centroid GPS coordinates of the most similar location for the available geographic information as calculated by the Nominatim search engine in OpenStreetMap. These columns need to use the column headers Continent, Country, State, Province, County, City, Area, Region, Location, and/or Site. EnteroBase provides these automatically but they would need to be manually included in the metadata table for the Standalone version of GrapeTree.
GrapeTree also transmits the color coding schemes and other metadata fields of all genomes in the visible tree to the MicroReact API interface, together with a Newick string representation of the tree itself, and opens the resulting MicroReact project in a separate browser window.
- References
- MicroReact: Argimon S, Abudahab K, Goater RJ, Fedosejev A, Bhai J, Glasner C, Feil EJ, Holden MT, Yeats CA, Grundmann H, et al. 2016. Microreact: visualizing and sharing data for genomic epidemiology and phylogeography. Microb Genom 2: e000093. URL: https://microreact.org
- OpenStreetMap: OpenStreetMap contributors. Planet dump retrieved from https://planet.osm.org. 2017. URL: https://www.openstreetmap.org
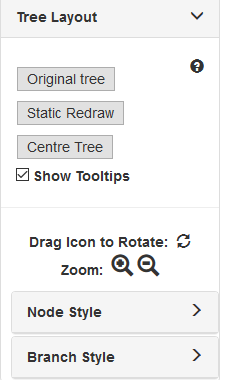
Tree Layout¶
- Original Tree (Standalone Only): Reverts the tree to the original state when it was loaded. You will lose all your changes!
- Static redraw: Redraws the tree using the static layout. You will lose any manual adjustments to node positioning!
- Centre Tree: Adjusts view settings to place entire tree in the centre of the window.
- Show Tooltips: Shows tooltips for branches and nodes.
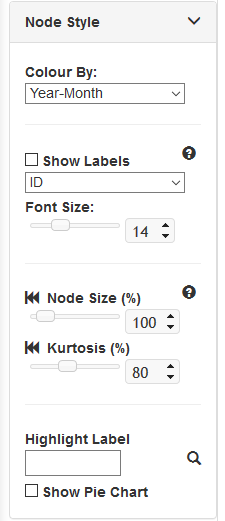
Node Style¶
- Show Labels: Check to show node labels and use dropdown to choose a label category.
- Font Size: Choose font size of node labels. Use the slider to change the value, or enter a specific value into the box
- Node Size: Increase/Decrease size of all nodes. Click rewind icon to revert to default value. Use the slider to change the value, or enter a specific value into the box
- Kurtosis: Increase/Decrease relative size of all nodes. Nodes with large number of members will look more distinct. Click rewind icon to revert to default value. Use the slider to change the value, or enter a specific value into the box
- Show Pie Chart: Shows one segment per genome within a node
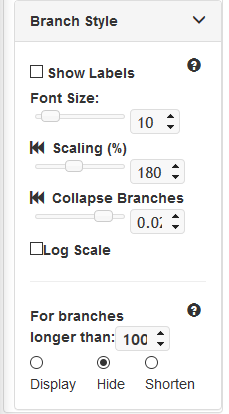
Branch Style¶
- Show Labels: Check to show node labels
- Font Size: Choose font size of node labels. Use the slider to change the value, or enter a specific value into the box.
- Scaling: Increase/Decrease length of all branches. Click rewind icon to revert to default value. Use the slider to change the value, or enter a specific value into the box.
- Collapse Branches: All branches shorter than specified length will be collapsed and nodes will merge as appropriate. Branch length value is scaled to the branch lengths defined in the original tree data. Use the slider to change the value, or enter a specific value into the box.
- Log Scale: All length of all branches will be scaled logarithmically.
Branches that are over a specified length can be rendered in a special manner according to the following settings in this panel.
- Display: Display all branches as normal
- Hide: Long branches will be invisible but remain interactive. Nodes will remain visible.
- Shorten: Long branches will be cropped back to the specified branch length cutoff. Lines will be dashed to indicate affected branches.
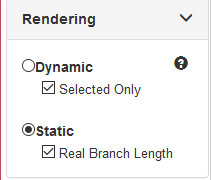
Layout Rendering options¶
Layout Rendering provides options for hot to positions nodes in the tree.
- Dynamic: Nodes are positioned dynamically similar to a Force Directed Layout. Nodes will try to fan out and distance themselves from neighbours. This may improve the aesthetics of the tree, but will modify branch length scaling. Branch lengths are NOT to scale when this is used. To only apply the dynamic positioning to selected nodes, check the “Selected Only” option.
- Static: Tree layout is calculated when the tree is initially created and remains static thereafter. Relative branch length scaling (as specified in the original tree data) will be maintained if “Real Branch Length” option is checked.
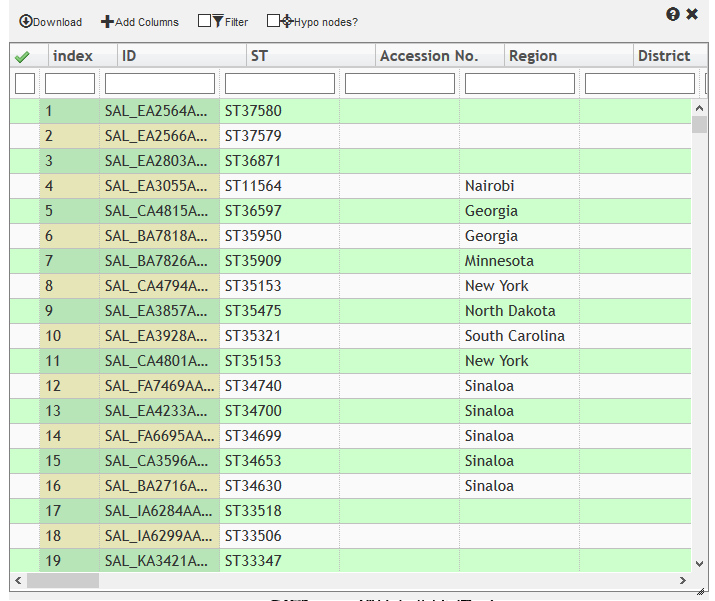
Metadata window¶
Provides a Slick-Grid (http://slickgrid.net/) editable table containing metadata. The first row shows Column headers. The second row allows filtering to show only those rows which contain the text in all of the text boxex. These text boxes also support regular expressions (engl|irel for England or Ireland; eng.*nd for England).
- Selected Column The leftmost column shows selected nodes as having a green check symbol. Clicking on the checkboxes in the left column also selects items in the tree. Selected items can be brought to the top by double-clicking on the
green arrow at the top left. It is possible to show only selected items by clccking the checkbox Selected Only in the top row, or by typing t (for True) in the left most checkbox under the green Selected Arrow. * Download: Export the metadata as a tab delimited file. * Add Metadata: Click to add a new column, and specify the field name in the column. In EnteroBase this offers the option to create a user-defined field in the main database. * Selected Only: Filter visible metadata to rows that are selected. Alternatively, type * Unfilter: Clear all filters permanently.